Estimating curvature measures from bivariate chi-bar-squared data using EM algorithm
estim_em_cm produces EM-type iterates from a two-column
matrix whose rows form iid samples from a bivariate chi-bar-squared
distribution.
estim_em_cm(d, low, upp, m_samp, N = 20, no_of_lcc_projections = 1, data = NULL)
Arguments
| d | the dimension of the bivariate chi-bar-squared distribution. |
|---|---|
| low | lower bound for |
| upp | upper bound for |
| m_samp | two-column matrix whose rows from iid samples from a bivariate chi-bar-squared distribution. |
| N | the number of iterates that shall be produced. |
| no_of_lcc_projections | number of projections on the log-concavity cone |
| data | output of |
Value
The output of estim_em_cm is a list of an (N+1)-by-(upp-low+1)
matrix whose rows constitute EM-type iterates, which may or may not
converge to the maximum likelihood estimate of the mixing weights of
the bivariate chi-bar-squared distribution, and the corresponding values
of the log-likelihood function.
Details
The sequence of iterates may or may not converge
to the maximum likelihood estimate of the mixing weights of the distribution.
Log-concavity of the weights is enforced by projecting the logarithms
onto the cone of log-concave sequences; this can be turned off by setting
no_of_lcc_projections=0.
This function is adapted from estim_em from the conivol
package, the difference being that the support of the weights is strictly
between the boundary cases. It is simplified in that the initial estimate
is always the uniform distribution, and the parity equation,
which does not hold for curvature measures, will not be enforced.
See also
estim_em,
constr_eigval,
constr_eigval_to_bcbsq,
prepare_em_cm,
indnorm_to_unnorm
Package: symconivol
Examples
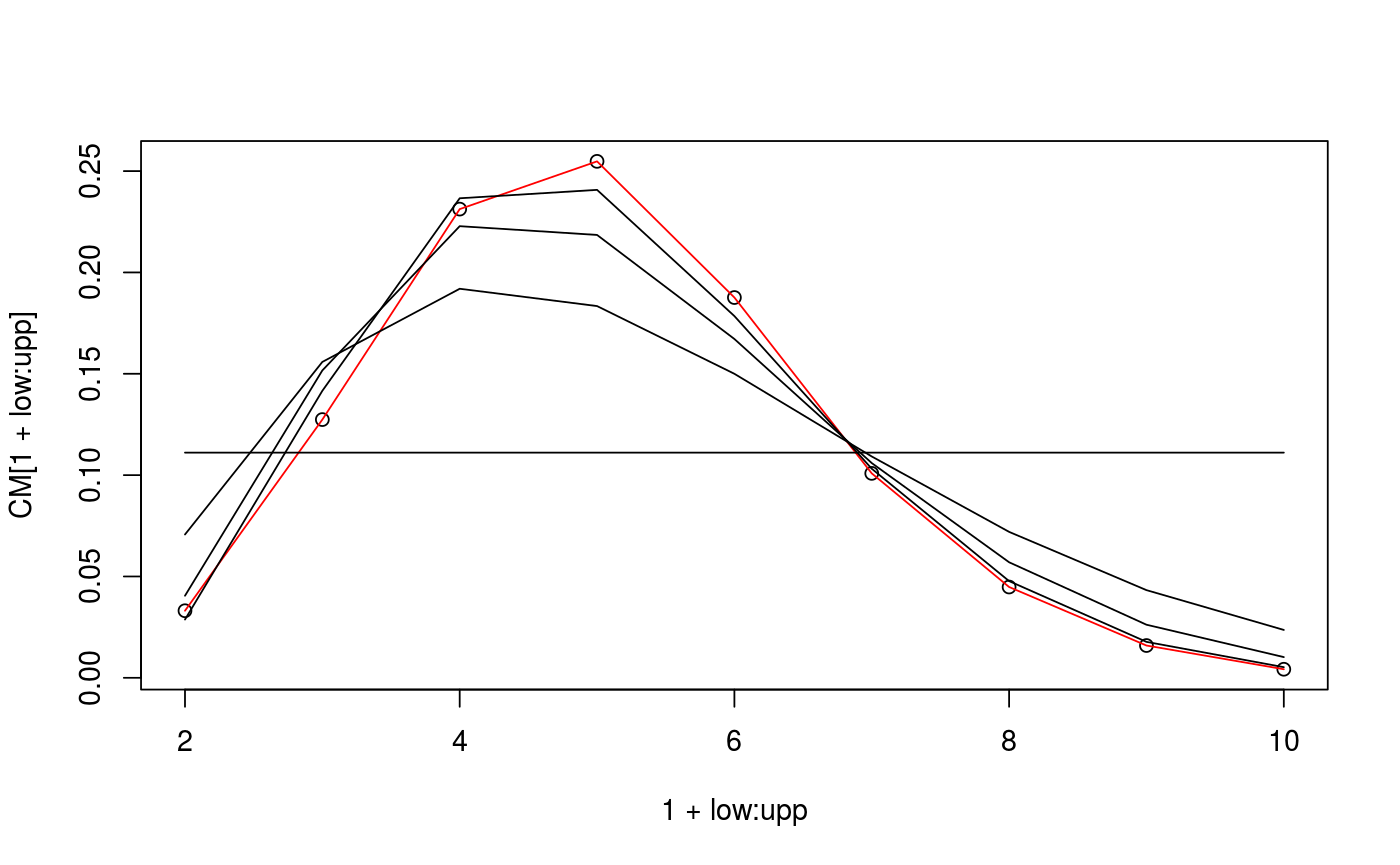
CM <- curv_meas_exact(4,3)$A[,2] CM <- CM/sum(CM) m_samp <- conivol::rbichibarsq(1e5,CM) d <- 15 low <- 1 upp <- 9 est <- estim_em_cm( d, low, upp, m_samp ) plot(1+low:upp, CM[1+low:upp])lines(1+low:upp, CM[1+low:upp], col="red")lines(1+low:upp, est[1,])lines(1+low:upp, est[5,])lines(1+low:upp, est[10,])lines(1+low:upp, est[21,])